论文原文:https://arxiv.org/pdf/1903.10122.pdf
Abstract
- Knowledge-driven Encode, Retrieve, Paraphrase (KERP) approach 知识驱动的编码、检索、释义(KERP)方法
- decomposes medical report generation into explicit medical abnormality graph learning 显式医学异常图学习 and subsequent natural language modeling 自然语言建模
visual features --(Encode Module)–> an abnormality graph --(Retireve module)–> sequences of templates --(Paraphrase module)–> sequences of words
- generates structured and robust reports supported with accurate abnormality prediction 生成结构化和健壮的报告,支持准确的异常预测
- produces explainable attentive regions which is crucial for interpretative diagnosis 产生可解释的注意区域,这对解释性诊断至关重要
GTR
-
core of KERP
-
dynamically transforms high-level semantics between graph-structured data of multiple domains such as knowledge graphs, images and sequences 在知识图、图像和序列等多个领域的图结构数据之间动态转换高级语义
GTR as a module
- concatenating intra-graph message passing and inter-graph message passing into one step 将图内消息传递和图间消息传递连接到一个步骤中
- conduct message passing within target graph 在目标图内进行消息传递
- conduct message passing from one / multiple source graph 从一个/多个源图传递消息
- stacking multiple such steps into one module 将多个这样的步骤叠加到一个模块中
- convert target graph features into high-level semantics 将目标图特性转换为高级语义
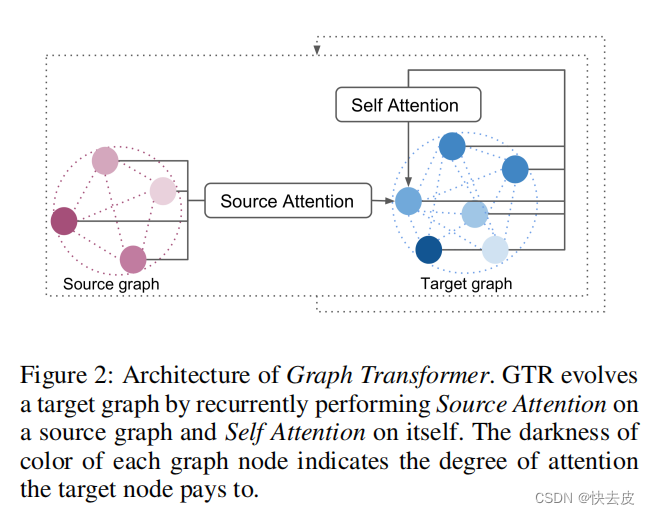
GTR as a multiple domains
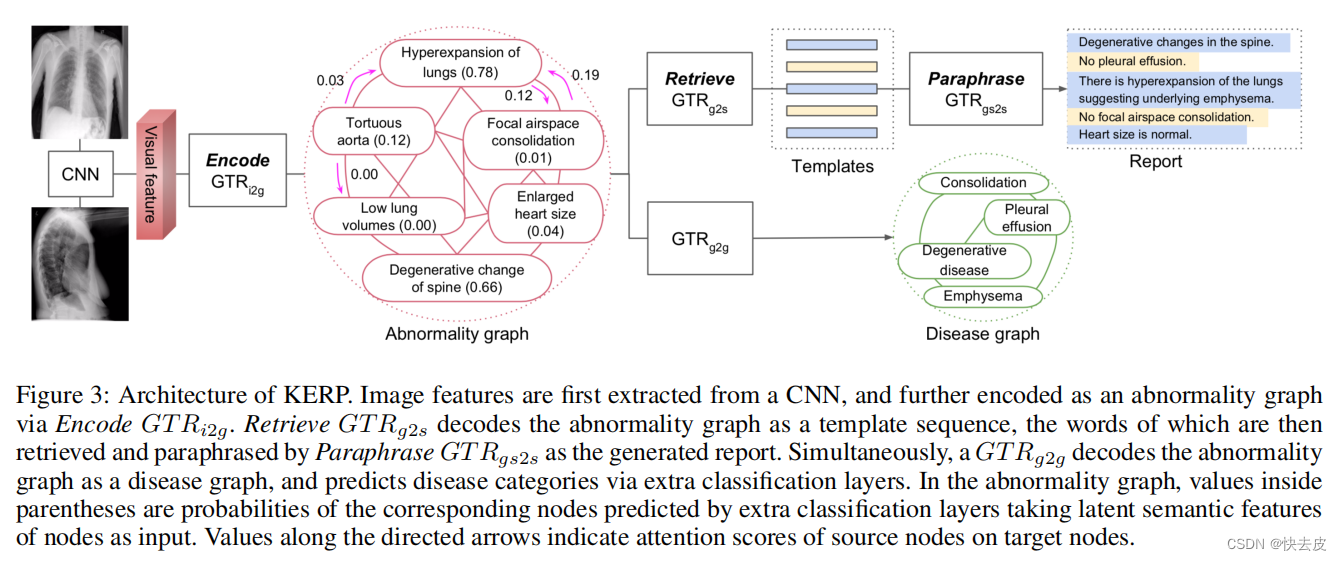
- GTR i2g \text{GTR}_\text{i2g} GTRi2g: image features --> graph’s features
- GTR g2s \text{GTR}_\text{g2s} GTRg2s: input - graph; output - sequence
- GTR g2g \text{GTR}_\text{g2g} GTRg2g: a graph --> another graph
- abnormality graph --> disease graph
- GTR gs2s \text{GTR}_\text{gs2s} GTRgs2s: input - graph&sequence; output - sequence
GTR for sequential input/output
positional encoding – relative and absolute position information 位置编码——相对和绝对位置信息
KERP
Encode module
- transforms visual features into a structured abnormality graph by incorporating prior medical knowledge 将视觉特征转化为结构化的异常图
- each node represents a possible clinical abnormality 临床异常
updated node features:
h u = G T R i 2 g ( X ) u = s i g m o i d ( W u h u ) \text{h}_u = GTR_{i2g}(\text{X}) \\ u = sigmoid(\text{W}_u\text{h}_u) hu=GTRi2g(X)u=sigmoid(Wuhu)
- W u \text{W}_u Wu: linear projection to transform latent feature u into 1-d probability 线性投影将潜在特征u转化为一维概率
- h u = ( h u 1 ; h u 2 ; . . . ; h u N ) ∈ R N , d \text{h}_u=(\text{h}_{u_1};\text{h}_{u_2};...;\text{h}_{u_N}) \in R^{N,d} hu=(hu1;hu2;...;huN)∈RN,d: the set of latent features of nodes where d is feature dimension 节点潜在特征集,其中d为特征维数
- u = ( u 1 , u 2 , . . . , u N ) , y i ∈ { 0 , 1 } , i ∈ { 1 , . . . , N } \text{u}=(u_1,u_2,...,u_N),y_i\in \{0,1\}, i\in\{1,...,N\} u=(u1,u2,...,uN),yi∈{ 0,1},i∈{ 1,...,N}:binary label for abnormality nodes 异常节点的二进制标签
Retrieve module 检索
- retrieves text templates based on the detected abnormalities 根据检测到的异常检索文本模板
obtain template sequence:
h t = G T R g 2 s ( h u ) t = argmax S o f t m a x ( W t h t ) \text{h}_t = GTR_{g2s}(\text{h}_u) \\ t = \text{argmax}Softmax(\text{W}_t\text{h}_t) ht=GTRg2s(hu)t=argmaxSoftmax(Wtht)
- W t \text{W}_t Wt: linear projection to transform latent feature to template embedding 线性投影 将潜在特征转化为模板嵌入
Paraphrase module
- refine templates with enriched details and possibly new case-specific findings 用丰富的细节和可能的新的特定病例发现来改进模板
- by modifying information in the templates that is not accurate for specific cases 通过修改模板中对于特定情况不准确的信息
- convert templates into more natural and dynamic expressions 将模板转换为更自然和生动的表达式
- by robust language modeling for the same content通过对同一内容进行稳健的语言建模
h w = G T R g s 2 s ( h u , t ) R = argmax S o f t m a x ( W w f ( h w ) ) \text{h}_w = GTR_{gs2s}(\text{h}_u,t) \\ R = \text{argmax}Softmax(\text{W}_wf(\text{h}_w)) hw=GTRgs2s(hu,t)R=argmaxSoftmax(Wwf(hw))
- f f f: the operation of reshaping h w \text{h}_w hw from R N s , N w , d R^{N_s,N_w,d} RNs,Nw,d to R N s ∗ N w , d R^{N_s*N_w,d} RNs∗Nw,d
- W w \text{W}_w Ww: linear projection to transform latent feature into word embedding 线性投影 将潜在特征转化为文字嵌入
Disease classification
multi-label disease classification: 多标记疾病分类
h z = G T R g 2 g ( h u ) z = s i g m o i d ( W z h z ) \text{h}_z = GTR_{g2g}(\text{h}_u) \\ z = sigmoid(\text{W}_z\text{h}_z) hz=GTRg2g(hu)z=sigmoid(Wzhz)
W z \text{W}_z Wz: linear projection to transform disease nodes feature into 1-d probability 线性投影 将疾病节点特征转化为一维概率
Learning
During paraphrasing, the retrieved templates t, instead of latent feature h t \text{h}_t ht, is used for rewriting. Sampling the templates of maximum predicted probability breaks the connectivity of differentiable back-propagation of the whole encode retrieve-paraphrase pipeline. 破坏了整个编码-检索-转述管道的可微调反向传播的连接性
- train the Paraphrase with ground truth templates
- then with **sampled templates **采样模板 generated by Retrieval module
Results

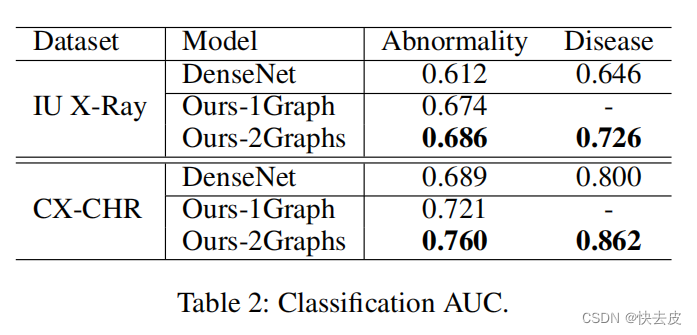
Conclusion
- accurate attributes prediction
- dynamic medical knowledge graph
- explainable location reference 可解释的位置参考