Multivariate Distributions
Copulas library supports several Multivariate distributions that support working with multiple random variables at the same time, taking into account the dependcies that my exist between them.
copulas.multivariate.GaussianMultivariate: Implements a multivariate distribution by combining the marginal univariate distributions with a Gaussian Copula.copulas.multivariate.VineCopula: Implements a multivariate distribution using Vine Copulas.
Gaussian Multivariate
from copulas.datasets import sample_trivariate_xyz
from copulas.visualization import scatter_3d
def func_Multivariate():
data = sample_trivariate_xyz()
print(data.head())
scatter_3d(data)
plt.show()
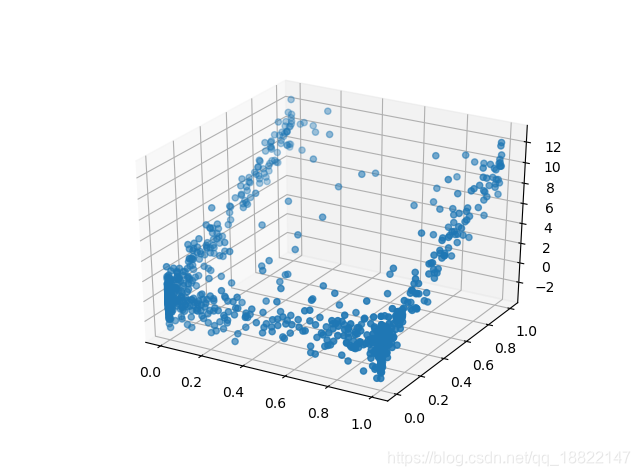
Fitting Model and Generating Synthetic Data by GaussianMultivariate class:
- Search for the
Univariatedistribution that better describes each column in thedata. - Fit the corresponding
Univariatedistributions to each column. - Learn the joint distribution based on the correlations between the marginal distributions
def func_Multivariate():
data = sample_trivariate_xyz()
print(data.head())
# scatter_3d(data)
# plt.show()
dist = GM()
dist.fit(data)
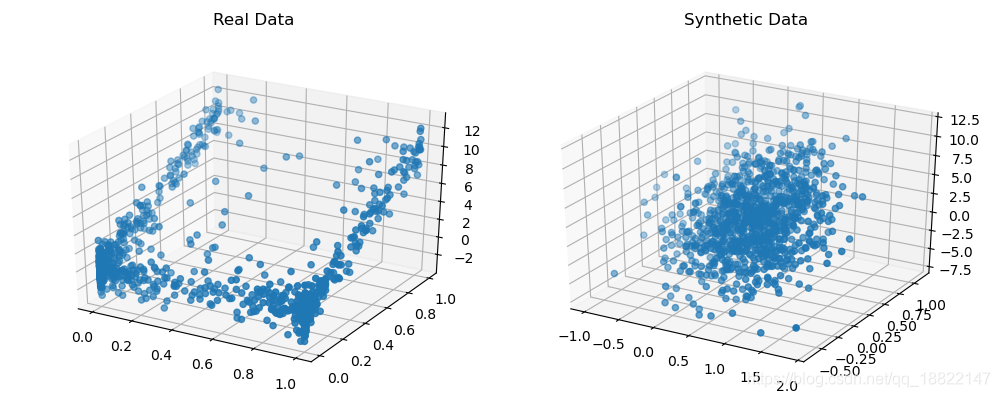
samples = dist.sample(1000)
compare_3d(data, samples)
plt.show()

Specifying column distributions. Advanced users can choose to manually specify the marginal distributions if they have additional information about the data.
dist = GaussianMultivariate(distribution=GaussianUnivariate)
dist.fit(data)
samples = dist.sample(1000)
compare_3d(data, samples)
plt.show()
By specifying the distribution that needs to be used in each column
dist = GaussianMultivariate(
distribution = {
'x': BetaUnivariate,
'y': GaussianKDE,
'z': GaussianUnivariate
})
dist.fit(data)
samples = dist.sample(1000)
compare_3d(data, samples)
plt.show()

By specifying a family of Univariate
univ = Univariate(parametric=ParametricType.PARAMETRIC)
dist = GaussianMultivariate(distribution=univ)
dist.fit(data)
samples = dist.sample(1000)
compare_3d(data, samples)
plt.show()

Vine Copulas
The Vine Copulas work by building a vine over the different columns in the dataset and estimating the pairwise (bivariate) relationship between the nodes on every edge.
def func_VineCopulas():
data = sample_trivariate_xyz()
center = VineCopula('center') # C-Copula
regular = VineCopula('regular') # R-Copula
direct = VineCopula('direct') # D-Copula
# Fitting data
center.fit(data)
regular.fit(data)
direct.fit(data)
c_samples = center.sample(1000)
r_samples = regular.sample(1000)
d_samples = direct.sample(1000)
compare_3d(data, c_samples)
plt.show()
