Johnson-Lindenstrauss开始嵌入随机投影
Johnson-Lindenstrauss引理指出,任何高维数据集都可以随机投影到较低维度的欧几里德空间,同时控制成对距离的失真。
理论界限
由随机投影p引入的失真由p定义具有良好概率的eps嵌入这一事实来确定:
(1−eps)‖u−v‖2<‖p(u)−p(v)‖2<(1+eps)‖u−v‖2
其中u和v是从形状[n_samples,n_features]的数据集中获取的任何行,而p是具有形状[n_components,n_features](或稀疏Achlioptas矩阵)的随机高斯N(0,1)矩阵的投影。
保证eps嵌入的最小组件数由下式给出:
n_components>=4log(n_samples)/(eps2/2−eps3/3)
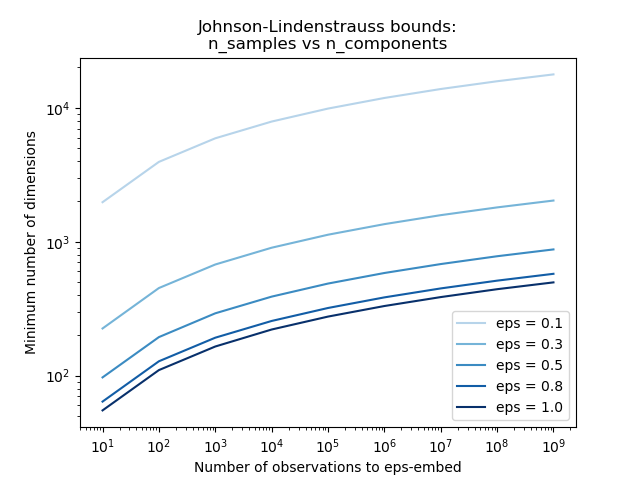
第一个图表显示,随着样本n_samples数量的增加,维数n_components的最小数量以对数方式增加,以保证eps嵌入。
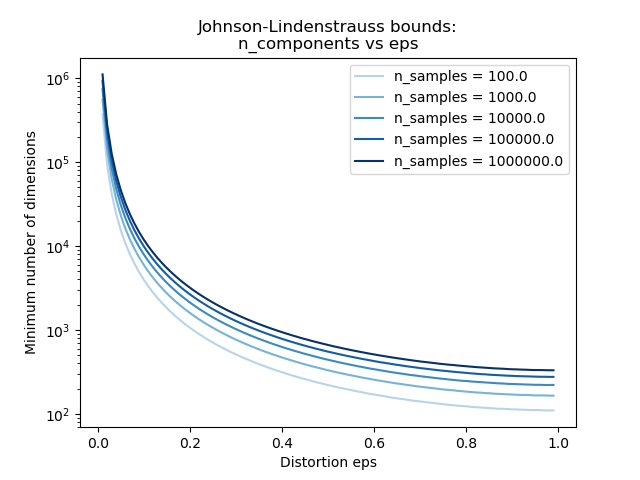
第二个图表显示,允许的失真eps的增加可以极大地减少给定数量的样本的n_components维数的最小数量n_samples
经验验证
我们在数字数据集或20个新闻组文本文档(TF-IDF词频)数据集上验证上述边界:
- 对于数字数据集,500个手写数字图片的一些8x8灰度级像素数据被随机投影到空间以用于各种更大数量的维度n_components。
- 对于20个新闻组数据集,使用稀疏随机矩阵将约500个具有100k特征的文档投影到较小的欧几里德空间,其具有针对目标维数n_components的不同值。
默认数据集是数字数据集。 要在二十个新闻组数据集上运行该示例,请将-twenty-newsgroups命令行参数传递给此脚本。
对于n_components的每个值,我们绘制:
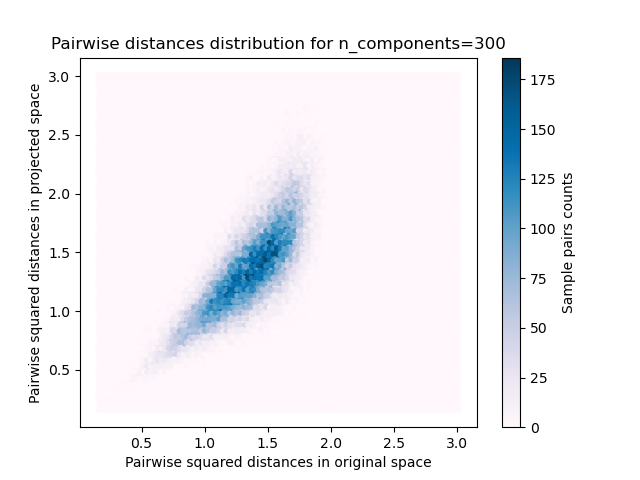
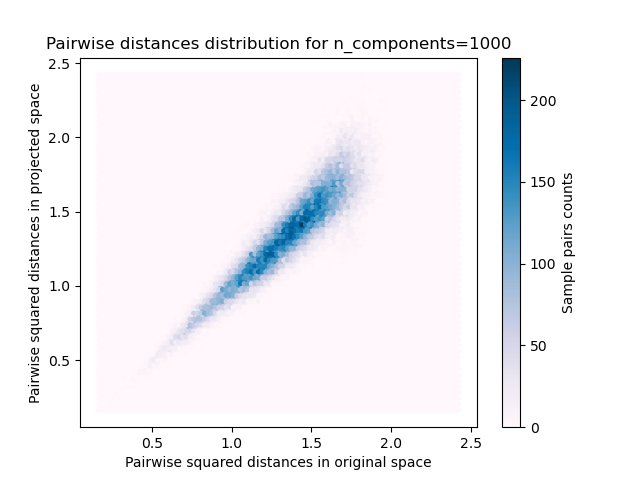
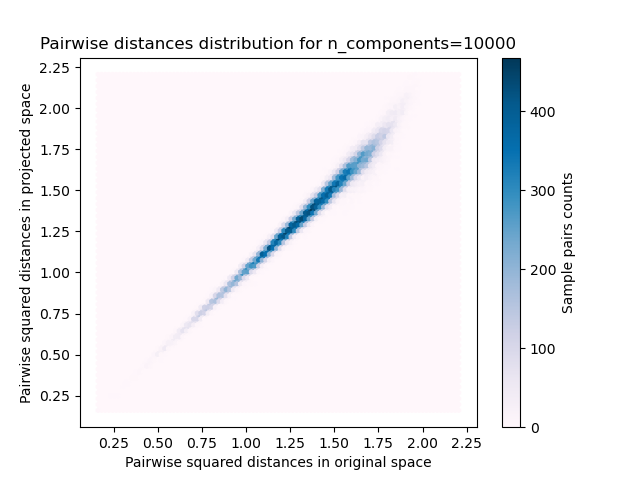
- 样本对的2D分布,原始和投影空间中的成对距离分别为x和y轴。
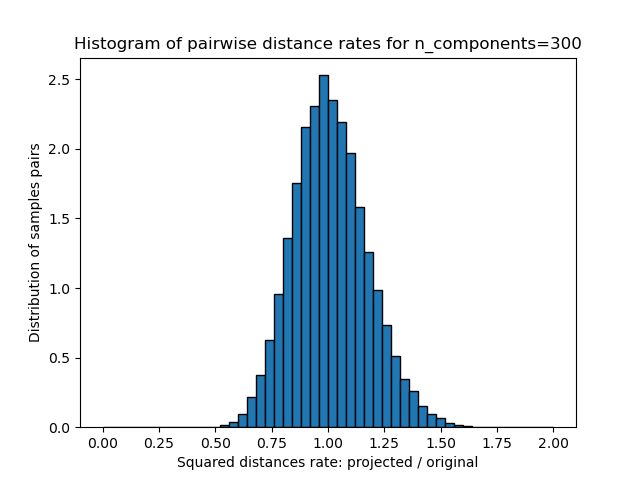
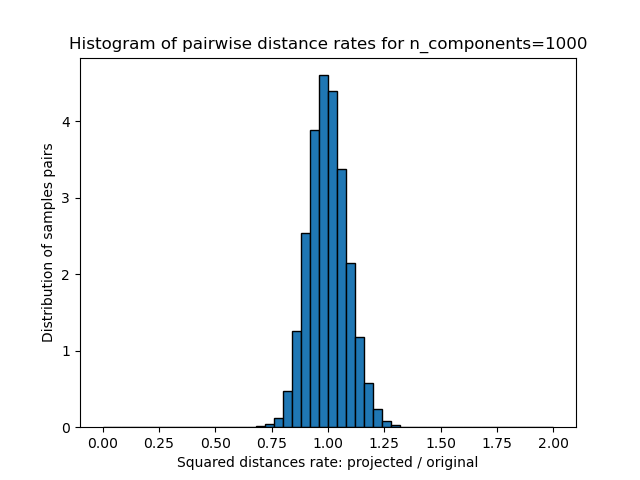
- 这些距离(投射/原始)的比率的1D直方图。
我们可以看到,对于n_components的低值,分布很宽,有许多失真的对和偏斜的分布(由于左边的零比率的硬限制,因为距离总是正的),而对于n_components的较大值,失真被控制并且 随机投影很好地保留了距离。
备注
根据JL引理,投影500个没有太多失真的样本将需要至少几千个维度,而不管原始数据集的特征数量。
因此,在输入空间中仅具有64个特征的数字数据集上使用随机投影是没有意义的:在这种情况下,它不允许降低维数。
另一方面,在二十个新闻组上,维度可以从56436降低到10000,同时合理地保留成对距离。
、
Out:
Embedding 500 samples with dim 64 using various random projections
Projected 500 samples from 64 to 300 in 0.005s
Random matrix with size: 0.029MB
Mean distances rate: 1.01 (0.07)
Projected 500 samples from 64 to 1000 in 0.015s
Random matrix with size: 0.096MB
Mean distances rate: 1.00 (0.05)
Projected 500 samples from 64 to 10000 in 0.215s
Random matrix with size: 0.963MB
Mean distances rate: 1.00 (0.02)print(__doc__)
import sys
from time import time
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from distutils.version import LooseVersion
from sklearn.random_projection import johnson_lindenstrauss_min_dim
from sklearn.random_projection import SparseRandomProjection
from sklearn.datasets import fetch_20newsgroups_vectorized
from sklearn.datasets import load_digits
from sklearn.metrics.pairwise import euclidean_distances
# `normed` is being deprecated in favor of `density` in histograms
if LooseVersion(matplotlib.__version__) >= '2.1':
density_param = {'density': True}
else:
density_param = {'normed': True}
# Part 1: plot the theoretical dependency between n_components_min and
# n_samples
# range of admissible distortions
eps_range = np.linspace(0.1, 0.99, 5)
colors = plt.cm.Blues(np.linspace(0.3, 1.0, len(eps_range)))
# range of number of samples (observation) to embed
n_samples_range = np.logspace(1, 9, 9)
plt.figure()
for eps, color in zip(eps_range, colors):
min_n_components = johnson_lindenstrauss_min_dim(n_samples_range, eps=eps)
plt.loglog(n_samples_range, min_n_components, color=color)
plt.legend(["eps = %0.1f" % eps for eps in eps_range], loc="lower right")
plt.xlabel("Number of observations to eps-embed")
plt.ylabel("Minimum number of dimensions")
plt.title("Johnson-Lindenstrauss bounds:\nn_samples vs n_components")
# range of admissible distortions
eps_range = np.linspace(0.01, 0.99, 100)
# range of number of samples (observation) to embed
n_samples_range = np.logspace(2, 6, 5)
colors = plt.cm.Blues(np.linspace(0.3, 1.0, len(n_samples_range)))
plt.figure()
for n_samples, color in zip(n_samples_range, colors):
min_n_components = johnson_lindenstrauss_min_dim(n_samples, eps=eps_range)
plt.semilogy(eps_range, min_n_components, color=color)
plt.legend(["n_samples = %d" % n for n in n_samples_range], loc="upper right")
plt.xlabel("Distortion eps")
plt.ylabel("Minimum number of dimensions")
plt.title("Johnson-Lindenstrauss bounds:\nn_components vs eps")
# Part 2: perform sparse random projection of some digits images which are
# quite low dimensional and dense or documents of the 20 newsgroups dataset
# which is both high dimensional and sparse
if '--twenty-newsgroups' in sys.argv:
# Need an internet connection hence not enabled by default
data = fetch_20newsgroups_vectorized().data[:500]
else:
data = load_digits().data[:500]
n_samples, n_features = data.shape
print("Embedding %d samples with dim %d using various random projections"
% (n_samples, n_features))
n_components_range = np.array([300, 1000, 10000])
dists = euclidean_distances(data, squared=True).ravel()
# select only non-identical samples pairs
nonzero = dists != 0
dists = dists[nonzero]
for n_components in n_components_range:
t0 = time()
rp = SparseRandomProjection(n_components=n_components)
projected_data = rp.fit_transform(data)
print("Projected %d samples from %d to %d in %0.3fs"
% (n_samples, n_features, n_components, time() - t0))
if hasattr(rp, 'components_'):
n_bytes = rp.components_.data.nbytes
n_bytes += rp.components_.indices.nbytes
print("Random matrix with size: %0.3fMB" % (n_bytes / 1e6))
projected_dists = euclidean_distances(
projected_data, squared=True).ravel()[nonzero]
plt.figure()
plt.hexbin(dists, projected_dists, gridsize=100, cmap=plt.cm.PuBu)
plt.xlabel("Pairwise squared distances in original space")
plt.ylabel("Pairwise squared distances in projected space")
plt.title("Pairwise distances distribution for n_components=%d" %
n_components)
cb = plt.colorbar()
cb.set_label('Sample pairs counts')
rates = projected_dists / dists
print("Mean distances rate: %0.2f (%0.2f)"
% (np.mean(rates), np.std(rates)))
plt.figure()
plt.hist(rates, bins=50, range=(0., 2.), edgecolor='k', **density_param)
plt.xlabel("Squared distances rate: projected / original")
plt.ylabel("Distribution of samples pairs")
plt.title("Histogram of pairwise distance rates for n_components=%d" %
n_components)
# TODO: compute the expected value of eps and add them to the previous plot
# as vertical lines / region
plt.show()