| 0 |
1 |
2 |
| 3 |
4 |
5 |
| 6 |
7 |
8 |
用前面一样的固定收敛标准,多次测量取平均值的办法比较不同的输入对迭代次数和分类准确率的影响。
一共设计了三组输入
| A |
B |
|||
| 0 |
4 |
8 |
< |
> |
| 0 |
4 |
|
< |
> |
| 0 |
8 |
< |
> |
|
| 4 |
8 |
< |
> |
|
| 0 |
< |
> |
||
| 1 |
< |
> |
||
| 2 |
< |
> |
||
| 3 |
< |
> |
||
| 4 |
< |
> |
||
| 5 |
< |
> |
||
| 6 |
< |
> |
||
| 7 |
< |
> |
||
| 8 |
< |
> |
||
| 9 |
< |
> |
(A,B)-9*9*2-(1,0)(0,1)
比如第一组输入A:让3*3矩阵的第0,4,8位为小于1的随机数,B为大于1的随机数,并用sigmoid函数处理。让A向(1,0)收敛,让B向(0,1)收敛。对应不同的收敛标准每个收敛标准重复199次取平均值和最大值。
得到的数据
| 048 |
04 |
08 |
48 |
0 |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
|
| δ |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
迭代次数n |
| 0.5 |
69.54774 |
87.04523 |
86.56281 |
85.39698 |
115.5377 |
114.8342 |
115.7387 |
115.9799 |
115.1859 |
115.2563 |
115.9095 |
115.2965 |
117.1859 |
| 0.4 |
4548.643 |
3712.528 |
3798.965 |
3726.985 |
3772.337 |
3785.884 |
3801.563 |
3814.442 |
3772.015 |
3781.317 |
3770.296 |
3788.045 |
3746.256 |
| 0.3 |
7019.281 |
5588.95 |
5556.266 |
5513.352 |
5343.683 |
5385.704 |
5320.035 |
5397.141 |
5361.864 |
5368.558 |
5328.528 |
5330.97 |
5292.749 |
| 0.2 |
10389.72 |
7897.462 |
7803.864 |
7875.191 |
7293.693 |
7368.106 |
7326.106 |
7295.02 |
7349.251 |
7265.834 |
7411.553 |
7299.704 |
7261.744 |
| 0.1 |
15640.72 |
12154.25 |
11925.91 |
11736.26 |
10720.67 |
10802.56 |
10842.2 |
10699.64 |
10881.95 |
10882.67 |
10788.34 |
10823.78 |
10856.22 |
| 0.01 |
37804.99 |
29799.56 |
29933.32 |
29667.67 |
26792.22 |
26901.37 |
26860.67 |
26813.95 |
27064.53 |
27012.71 |
26750.53 |
27287.55 |
26885.66 |
| 0.001 |
70411.91 |
60322.03 |
58871.21 |
60065.49 |
55179.8 |
54979.79 |
55442.9 |
55365.78 |
55606.25 |
55015.88 |
55099.73 |
55721.57 |
55648.81 |
| 1.00E-04 |
120662.3 |
105158.8 |
106917.8 |
106076.2 |
100157.3 |
103572.5 |
102357 |
101774.3 |
101938.9 |
101639.3 |
102363.5 |
101993.4 |
102015.4 |
| 9.00E-05 |
121179 |
108435.7 |
108578.8 |
109015.9 |
105790.8 |
104659.5 |
104810.9 |
105340.9 |
103461.6 |
104283.4 |
105418.5 |
103671.1 |
105346.9 |
| 8.00E-05 |
130110.8 |
111452 |
113190.3 |
113299.7 |
107869.5 |
108540.3 |
108181.4 |
105928.8 |
107891.4 |
107789.7 |
108550.8 |
106467.5 |
108052.4 |
| 7.00E-05 |
126860.3 |
115019 |
115761.6 |
115877.4 |
111307.1 |
110133 |
110952.4 |
111819.5 |
111779.4 |
111654.9 |
111009.9 |
111891.3 |
110574.1 |
| 6.00E-05 |
132585.4 |
119918.6 |
119752.3 |
121557.7 |
115674.8 |
117992.8 |
115647.7 |
117834.6 |
117606.9 |
116615.9 |
115671.1 |
115095.3 |
115095.1 |
| 5.00E-05 |
137890.7 |
125209 |
125577.9 |
125954 |
120692.8 |
119851.7 |
119278 |
119960.9 |
118475.3 |
120766.5 |
119895.6 |
120370.2 |
120516.1 |
| 4.00E-05 |
147064.4 |
133217.3 |
131870.8 |
129671.7 |
127405.8 |
127971.9 |
129132.7 |
129201.6 |
127790.6 |
127458.8 |
128005.5 |
128684.2 |
127511 |
| 3.00E-05 |
155796.4 |
139982.7 |
141179.5 |
140967.1 |
136719.7 |
134369.5 |
140037.1 |
135600 |
138670.1 |
135805.1 |
134472.2 |
137149.9 |
137316.7 |
| 2.00E-05 |
169985.5 |
154108.3 |
154507.8 |
153714.9 |
150518.4 |
151828.2 |
148436.7 |
151129.8 |
149605.7 |
150349.2 |
150842.8 |
150540.6 |
150175.1 |
| 1.00E-05 |
197630.1 |
179095.2 |
177795.9 |
179693.8 |
174674.7 |
176821.8 |
176704.8 |
175695.2 |
177221.7 |
177198.3 |
176186.3 |
175188.5 |
176742.3 |
| 9.00E-06 |
196676.7 |
180193.5 |
182343.7 |
178640.7 |
179333.3 |
181265.5 |
179465.4 |
182904.4 |
179777.1 |
180433.6 |
180775.9 |
177657.7 |
182068.1 |
| 8.00E-06 |
199874.4 |
186751.9 |
188153.9 |
183630.4 |
183541.9 |
188428.1 |
186458.7 |
187255.6 |
186014.6 |
184091.9 |
185337.7 |
185151.7 |
187143.4 |
| 7.00E-06 |
216151.2 |
193697.2 |
192957.5 |
194736.5 |
192517.3 |
192300.2 |
193270.8 |
192667.4 |
192103.2 |
191011.7 |
192621.6 |
190140.4 |
189724.4 |
| 6.00E-06 |
217972.8 |
200086.2 |
202795.4 |
200274.7 |
195581.3 |
203314.2 |
196539.2 |
197712.5 |
199183.8 |
197640.4 |
199489.3 |
197581.3 |
198676.2 |
| 5.00E-06 |
223973.4 |
203460.9 |
208344.9 |
209322.7 |
203278.2 |
204251.4 |
203931.9 |
209619.4 |
205602.9 |
205946.4 |
203393.3 |
205240.8 |
206051.4 |
| 4.00E-06 |
241939.3 |
217785 |
209999.1 |
211848.5 |
212986.2 |
215625.3 |
216182.7 |
217577.8 |
215103.6 |
216877.4 |
214410.2 |
215425.9 |
217087.1 |
| 3.00E-06 |
251392.4 |
230528.7 |
228678.2 |
232201.8 |
229598.3 |
227561.4 |
233740.7 |
229803 |
233331.3 |
226590.8 |
228951.3 |
231160.4 |
230334.5 |
| 2.00E-06 |
278920.6 |
251832.5 |
255143.9 |
255012.3 |
251758.4 |
254739.4 |
252221.8 |
254074.7 |
248451.8 |
250446.2 |
251003.4 |
247446.5 |
250214.4 |
| 1.00E-06 |
320757.6 |
283314.5 |
284455.6 |
288156 |
294319.3 |
294355.5 |
296194.7 |
287404.2 |
296014.9 |
290935.4 |
294212 |
292735.3 |
290289.1 |
| 143588.8 |
129184.9 |
129460.8 |
129550.9 |
127036.3 |
127958.5 |
127817.5 |
127800.2 |
127699.1 |
127191.4 |
127379.8 |
127075.3 |
127490.1 |
得到结果很明显当输入位只有1个的时候迭代次数是大约相同的,当输入位只有两个的时候得到的迭代次数也大约是相同的。

比较平均值
![]()
按照相同收敛标准下迭代次数越大两个被分类对象之间的差异越小的方法比较
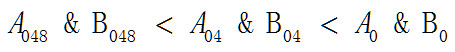
矩阵A048和矩阵B048之间的差异要小于矩阵A04和B04之间的差异,矩阵A04和B04之间的差异要小于矩阵A0和B0之间的差异。这个结果表明两个分类对象在位置固定的前提下,差异的大小只与数值有关而与具体位置无关。
| 048 |
04 |
08 |
48 |
0 |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
|
| δ |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
平均准确率p-ave |
| 0.5 |
0.500571 |
0.501507 |
0.499077 |
0.500214 |
0.500747 |
0.501502 |
0.500737 |
0.500601 |
0.499967 |
0.500179 |
0.502608 |
0.499339 |
0.499288 |
| 0.4 |
0.978521 |
0.96493 |
0.963521 |
0.963733 |
0.923889 |
0.927591 |
0.921741 |
0.925458 |
0.928119 |
0.92577 |
0.923491 |
0.926645 |
0.925569 |
| 0.3 |
0.995775 |
0.985382 |
0.985966 |
0.985392 |
0.956962 |
0.955393 |
0.95417 |
0.949331 |
0.953089 |
0.954839 |
0.95239 |
0.953838 |
0.952485 |
| 0.2 |
0.997877 |
0.992243 |
0.992495 |
0.99167 |
0.970211 |
0.973878 |
0.970357 |
0.971982 |
0.97416 |
0.975312 |
0.972435 |
0.970921 |
0.971871 |
| 0.1 |
0.999281 |
0.996504 |
0.996811 |
0.996826 |
0.986514 |
0.984467 |
0.984039 |
0.984482 |
0.985719 |
0.985186 |
0.984678 |
0.98578 |
0.985669 |
| 0.01 |
0.966394 |
0.989512 |
0.989723 |
0.989562 |
0.958637 |
0.960493 |
0.9591 |
0.959693 |
0.961383 |
0.958924 |
0.961655 |
0.960282 |
0.961897 |
| 0.001 |
0.827159 |
0.957752 |
0.960086 |
0.960061 |
0.928099 |
0.926887 |
0.929266 |
0.929326 |
0.928195 |
0.926675 |
0.928848 |
0.930503 |
0.925468 |
| 1.00E-04 |
0.69811 |
0.914055 |
0.910493 |
0.909462 |
0.911922 |
0.907536 |
0.907993 |
0.906167 |
0.908446 |
0.909296 |
0.909015 |
0.905936 |
0.90916 |
| 9.00E-05 |
0.672039 |
0.909477 |
0.908693 |
0.900765 |
0.907375 |
0.905669 |
0.908778 |
0.908009 |
0.908169 |
0.908622 |
0.910529 |
0.907309 |
0.906323 |
| 8.00E-05 |
0.675148 |
0.912254 |
0.902838 |
0.906248 |
0.906173 |
0.904241 |
0.911323 |
0.907727 |
0.908396 |
0.908245 |
0.906882 |
0.908421 |
0.906625 |
| 7.00E-05 |
0.678206 |
0.906776 |
0.901842 |
0.904085 |
0.908552 |
0.907707 |
0.90664 |
0.90824 |
0.906907 |
0.908491 |
0.906328 |
0.90905 |
0.907762 |
| 6.00E-05 |
0.653281 |
0.895212 |
0.896489 |
0.892521 |
0.90748 |
0.906152 |
0.906097 |
0.904814 |
0.90577 |
0.906323 |
0.908959 |
0.906283 |
0.905483 |
| 5.00E-05 |
0.653256 |
0.892888 |
0.88907 |
0.889447 |
0.904955 |
0.906419 |
0.904145 |
0.90736 |
0.904301 |
0.905509 |
0.906067 |
0.905846 |
0.906771 |
| 4.00E-05 |
0.643548 |
0.883642 |
0.887239 |
0.888713 |
0.903637 |
0.901967 |
0.904482 |
0.901997 |
0.907023 |
0.903134 |
0.907807 |
0.906434 |
0.904845 |
| 3.00E-05 |
0.644051 |
0.873572 |
0.876354 |
0.879809 |
0.905544 |
0.905941 |
0.90152 |
0.90416 |
0.899543 |
0.904799 |
0.907259 |
0.902616 |
0.902953 |
| 2.00E-05 |
0.621778 |
0.868134 |
0.868004 |
0.870418 |
0.903225 |
0.901766 |
0.904437 |
0.905398 |
0.903894 |
0.902174 |
0.901027 |
0.900685 |
0.900861 |
| 1.00E-05 |
0.602261 |
0.852461 |
0.85317 |
0.854317 |
0.89987 |
0.899588 |
0.901972 |
0.902229 |
0.902008 |
0.901323 |
0.898688 |
0.900715 |
0.895825 |
| 9.00E-06 |
0.605671 |
0.857546 |
0.857979 |
0.856761 |
0.900066 |
0.901082 |
0.901977 |
0.900021 |
0.899885 |
0.898793 |
0.901007 |
0.902078 |
0.900438 |
| 8.00E-06 |
0.597225 |
0.851983 |
0.846932 |
0.855303 |
0.901454 |
0.899281 |
0.898019 |
0.900237 |
0.898813 |
0.899819 |
0.899669 |
0.901756 |
0.900861 |
| 7.00E-06 |
0.601189 |
0.843592 |
0.847249 |
0.840071 |
0.899955 |
0.901203 |
0.900554 |
0.899176 |
0.900156 |
0.900438 |
0.897888 |
0.900363 |
0.900866 |
| 6.00E-06 |
0.609499 |
0.838577 |
0.833517 |
0.845619 |
0.900197 |
0.897561 |
0.90078 |
0.902058 |
0.899578 |
0.900146 |
0.898114 |
0.900664 |
0.898612 |
| 5.00E-06 |
0.588694 |
0.845881 |
0.835252 |
0.84078 |
0.89993 |
0.899754 |
0.89989 |
0.899226 |
0.899558 |
0.900141 |
0.899719 |
0.89997 |
0.900222 |
| 4.00E-06 |
0.588644 |
0.839453 |
0.84411 |
0.844624 |
0.900187 |
0.900836 |
0.898793 |
0.899875 |
0.899583 |
0.898668 |
0.898411 |
0.900141 |
0.899744 |
| 3.00E-06 |
0.581692 |
0.829669 |
0.838688 |
0.830449 |
0.896726 |
0.896339 |
0.898139 |
0.899663 |
0.897008 |
0.898476 |
0.8966 |
0.901273 |
0.898708 |
| 2.00E-06 |
0.589293 |
0.823668 |
0.813603 |
0.81654 |
0.899397 |
0.899915 |
0.899045 |
0.897174 |
0.900539 |
0.898904 |
0.897365 |
0.898562 |
0.896922 |
| 1.00E-06 |
0.569399 |
0.816777 |
0.810927 |
0.808024 |
0.898849 |
0.896821 |
0.899497 |
0.897189 |
0.899628 |
0.896484 |
0.898607 |
0.89902 |
0.897068 |
| 0.697637 |
0.878594 |
0.877313 |
0.877747 |
0.899252 |
0.898846 |
0.898981 |
0.898907 |
0.899225 |
0.899103 |
0.899079 |
0.899401 |
0.89855 |
相当奇怪的是随着被分类元素增加分类准确率是下降的
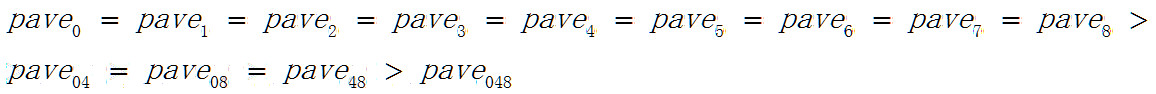
矩阵A048可以看成是由矩阵A04,A08,A48组成的,但A048和B048的分类准确率确下降了。
比较最大分类准确率
| 048 |
04 |
08 |
48 |
0 |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
|
| δ |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
最大准确率p-max |
| 0.5 |
0.550551 |
0.553554 |
0.542543 |
0.545546 |
0.540541 |
0.613614 |
0.65966 |
0.626627 |
0.556557 |
0.555556 |
0.542543 |
0.534535 |
0.536537 |
| 0.4 |
0.997998 |
0.994995 |
0.994995 |
0.993994 |
0.973974 |
0.97998 |
0.975976 |
0.977978 |
0.996997 |
0.983984 |
0.975976 |
0.977978 |
0.977978 |
| 0.3 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0.991992 |
1 |
1 |
1 |
1 |
1 |
| 0.2 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
| 0.1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
| 0.01 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
| 0.001 |
1 |
1 |
1 |
1 |
0.980981 |
0.968969 |
0.980981 |
0.97998 |
0.977978 |
0.977978 |
0.976977 |
0.97998 |
0.978979 |
| 1.00E-04 |
0.997998 |
0.995996 |
0.996997 |
0.98999 |
0.968969 |
0.958959 |
0.958959 |
0.948949 |
0.951952 |
0.961962 |
0.950951 |
0.961962 |
0.956957 |
| 9.00E-05 |
0.973974 |
0.993994 |
0.996997 |
0.992993 |
0.953954 |
0.947948 |
0.95996 |
0.955956 |
0.960961 |
0.975976 |
0.955956 |
0.955956 |
0.966967 |
| 8.00E-05 |
0.96997 |
0.991992 |
0.986987 |
0.992993 |
0.948949 |
0.963964 |
0.954955 |
0.952953 |
0.958959 |
0.945946 |
0.952953 |
0.952953 |
0.946947 |
| 7.00E-05 |
0.997998 |
0.990991 |
0.990991 |
0.993994 |
0.956957 |
0.962963 |
0.951952 |
0.954955 |
0.955956 |
0.958959 |
0.950951 |
0.954955 |
0.951952 |
| 6.00E-05 |
0.98999 |
0.990991 |
0.990991 |
0.987988 |
0.964965 |
0.971972 |
0.950951 |
0.962963 |
0.954955 |
0.952953 |
0.955956 |
0.948949 |
0.94995 |
| 5.00E-05 |
0.984985 |
0.982983 |
0.990991 |
0.997998 |
0.954955 |
0.94995 |
0.956957 |
0.94995 |
0.953954 |
0.955956 |
0.955956 |
0.962963 |
0.95996 |
| 4.00E-05 |
0.93994 |
0.988989 |
0.990991 |
0.991992 |
0.954955 |
0.942943 |
0.95996 |
0.955956 |
0.951952 |
0.966967 |
0.95996 |
0.945946 |
0.952953 |
| 3.00E-05 |
0.992993 |
0.985986 |
0.987988 |
0.98999 |
0.952953 |
0.950951 |
0.945946 |
0.961962 |
0.943944 |
0.953954 |
0.950951 |
0.946947 |
0.954955 |
| 2.00E-05 |
0.974975 |
0.986987 |
0.991992 |
0.990991 |
0.947948 |
0.946947 |
0.94995 |
0.951952 |
0.951952 |
0.953954 |
0.947948 |
0.946947 |
0.946947 |
| 1.00E-05 |
0.908909 |
0.976977 |
0.991992 |
0.987988 |
0.946947 |
0.945946 |
0.955956 |
0.938939 |
0.938939 |
0.954955 |
0.943944 |
0.944945 |
0.94995 |
| 9.00E-06 |
0.956957 |
0.984985 |
0.991992 |
0.971972 |
0.942943 |
0.94995 |
0.948949 |
0.951952 |
0.942943 |
0.945946 |
0.938939 |
0.937938 |
0.935936 |
| 8.00E-06 |
0.900901 |
0.994995 |
0.986987 |
0.978979 |
0.953954 |
0.940941 |
0.946947 |
0.943944 |
0.935936 |
0.95996 |
0.950951 |
0.944945 |
0.947948 |
| 7.00E-06 |
0.965966 |
0.980981 |
0.968969 |
0.970971 |
0.941942 |
0.950951 |
0.942943 |
0.94995 |
0.94995 |
0.943944 |
0.944945 |
0.946947 |
0.944945 |
| 6.00E-06 |
0.928929 |
0.970971 |
0.978979 |
0.96997 |
0.942943 |
0.941942 |
0.94995 |
0.951952 |
0.937938 |
0.944945 |
0.947948 |
0.947948 |
0.938939 |
| 5.00E-06 |
0.963964 |
0.973974 |
0.96997 |
0.97998 |
0.941942 |
0.935936 |
0.93994 |
0.93994 |
0.947948 |
0.943944 |
0.934935 |
0.941942 |
0.957958 |
| 4.00E-06 |
0.862863 |
0.966967 |
0.965966 |
0.977978 |
0.947948 |
0.941942 |
0.933934 |
0.943944 |
0.93994 |
0.950951 |
0.93994 |
0.947948 |
0.94995 |
| 3.00E-06 |
0.8999 |
0.973974 |
0.986987 |
0.957958 |
0.948949 |
0.936937 |
0.936937 |
0.940941 |
0.935936 |
0.941942 |
0.963964 |
0.940941 |
0.940941 |
| 2.00E-06 |
0.918919 |
0.980981 |
0.993994 |
0.984985 |
0.944945 |
0.940941 |
0.93994 |
0.938939 |
0.933934 |
0.935936 |
0.938939 |
0.938939 |
0.941942 |
| 1.00E-06 |
0.855856 |
0.967968 |
0.964965 |
0.978979 |
0.943944 |
0.937938 |
0.938939 |
0.930931 |
0.941942 |
0.937938 |
0.942943 |
0.935936 |
0.934935 |
|
|
0.943636 |
0.970393 |
0.971664 |
0.970316 |
0.944483 |
0.945484 |
0.947717 |
0.946292 |
0.943135 |
0.946331 |
0.943251 |
0.94225 |
0.943251 |
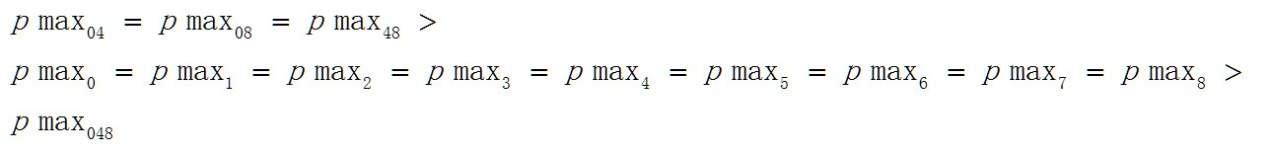
Pmax04>pmax0>pmax048,这一顺序与平均值的顺序并不相同。
可以很容易用物理上对称性破缺的概念来理解神经网络的分类行为,比如如果两个对象可以通过一个网络实现分类,表明这两个对象相对这个神经网络是对称性破缺的。如果两个对象无法通过某个神经网络实现分类,也就是分类准确率恒为0.5,也就表明这两个对象相对这个神经网络是对称的。有此可以认为分类准确率反映了两个分类对象相对神经网络的对称性。
参照平均分类准确率pave可以得出随着分类元素的增加,分类对象之间的对称性是增加的。
